Hi,
Making alignments of RNAs reveals conservation and thereby possible
important regions (think of guide-RNA sections of snoRNAs or
protein-binding regions such as the box C/D regions). This can be
visualized with the WEB-logo / consensus below the sequence by:
a) tick "autocalculate consensus" under "Calculate"
b) tick "show annotations" under "Annotations"
c) tick "show consensus logo" in "autocalculated annotation" under
"Annotation"
Then, by linking up with RNAalifold, that is
d) tick "RNAalifold Prediction" in "Secondary Structure prediction"
under "Web Service",
you also get common secondary structure elements shown below the
alignment. By right-clicking on the labels in the margin each
annotation by itself can be viewed/saved.
All can be saved in the .jvp jalview project-file format, incl. format
- and colour settings.
This is great.
But I experience some serious limitations that prevent me to make
more use of all this info. Maybe I missed some ways to do this so any
info to achieve this will be great.
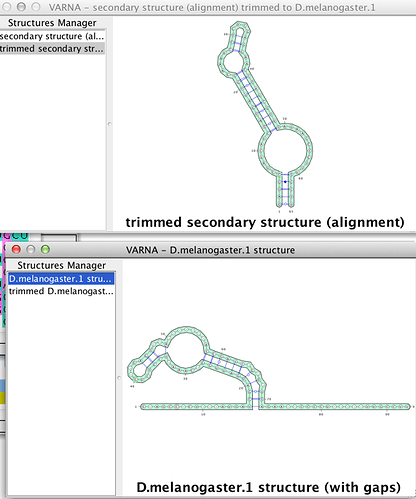
For example, I cannot view the generated 2D-structurethis directly in
VARNA via Jalview (only sequences loaded with structure information
seems to trigger this option, not sequences with structure information
added inside the program).
The generated WebLogo can be saved as part of the complete
alignment but not as an image by itself (which I would find the most
useful aspect of having this logo as it abbreviates a space-consuming
alignment and would bypass the need to do make such a figure on the
WebLogo site).
Also, the MFE-structure generated by alifold is sometimes not complete
but manual editing is not possible nor can the dot-bracket line be
copied for pasting into VARNA manually.
This limitation I bypass by generating a fake-sequence using
import-from text-box in which I use dot-bracket notation to indicate
secondary structure elements;
structure
.......((((....))))...........
That line, together with a reference sequence I copy manually
into VARNA and use it to draw a 2D-structure model. But each edit/change
will require repeating this copy/pasting manually. VARNA itself makes
this editing also quite tedious (each edit generates a new structure
that has to be reformatted).
The biggest lost though, is the conservation-information that is
embedded in the consensus and web-logo. VARNA allows colouration of
nucleotides by using a Color Map. But I cannot find an easy way of
converting the Consensus-WebLogo csv file to a version that could be
used as a colormap in Varna.
A way around this would possibly be to:
a) create a 'structure field/line' that is read by
JalView as such (i.e. that triggers the connection with VARNA) in which
an alifold output can be loaded but that is also editable. Ideally this
field would allow saving to various RNA-2D formats (bracket,
stockholm,ct)
b) create a way to export the weblogo-conservation info to a color-map
for VARNA (or load this into the VARNA displayed structure).
Does this sound feasible or are other tools at hand I have missed to
accomplish this???
Cheers,
Rob
···
--
Dr. R.W. van Nues
SYNTHETIC and SYSTEMS BIOLOGY at EDINBURGH (SynthSys)
The University of Edinburgh
CH Waddington Building
Max Born Crescent
Edinburgh EH9 3BF
Telephone +44 (0)131 651 9018
E-mail:rob.van-nues@ed.ac.uk
The University of Edinburgh is a charitable body, registered in
Scotland, with registration number SC005336.